Jurisdiction:
Sweden
Organ System:
Skin
Funding Organizations:
- Fru Berta Kamprads Stiftelse
- Mats and Stefan Paulsson Trust
- Conselho Nacional de Desenvolvimento Científico e Tecnológico
- Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro
- Coordenação de Aperfeiçoamento de Pessoal de Nível Superior
- Hungarian Academy of Sciences
- Országos Tudományos Kutatási Alapprogramok
- National Research Foundation of Korea
Research Organizations:
- Lund University, Sweden
- Semmelweis University, Hungary
- Karolinska Institute, Sweden
- Federal University of Rio de Janeiro, Brazil
Principal Investigators
:- István Balázs Németh
- György Marko-Varga
Publications:
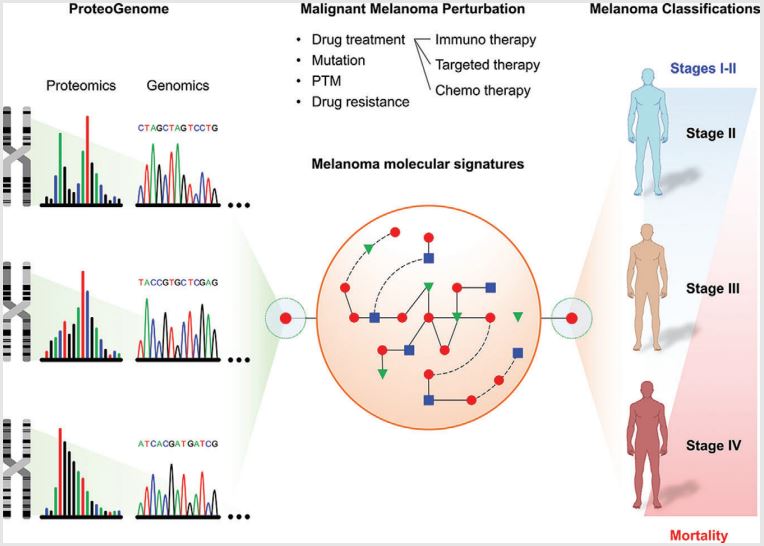
First study is to map the protein levels in malignant melanoma tumor samples, focused on in-depth histopathology coupled to proteome characterization. The protein levels and localization were determined for a broad spectrum of diverse, surgically isolated melanoma tumors originating from multiple body locations. More than 15,500 proteoforms were identified by mass spectrometry, from which chromosomal and subcellular localization was annotated within both primary and metastatic melanoma. The data generated by global proteomic experiments covered 72% of the proteins identified in the recently reported high stringency blueprint of the human proteome. This study contributes to the NIH Cancer Moonshot initiative combining detailed histopathological presentation with the molecular characterization for 505 melanoma tumor samples, localized in 26 organs from 232 patients.
The second study is to establish a knowledge basis of the tumor proteome to serve as a complement to genome and transcriptome studies. The melanoma proteome landscape, obtained by the analysis of 505 well-annotated melanoma tumor samples, is defined based on almost 16 000 proteins, including mutated proteoforms of driver genes. This data covers 65% and 74% of the predicted and identified human proteome, respectively.
The data that support the findings of both studies are openly available in ProteomeXchange at http://www.proteomexchange.org/, reference numbers PXD001725, PXD001724, PXD009630, PXD017968, and PXD026086